SARS-CoV-2 Spike RBD Recombinant Nanobody
-
中文名稱:SARS-CoV-2 Spike RBD重組納米抗體
-
貨號:CSB-RA33245A2GMY
-
規(guī)格:¥3080
-
圖片:
-
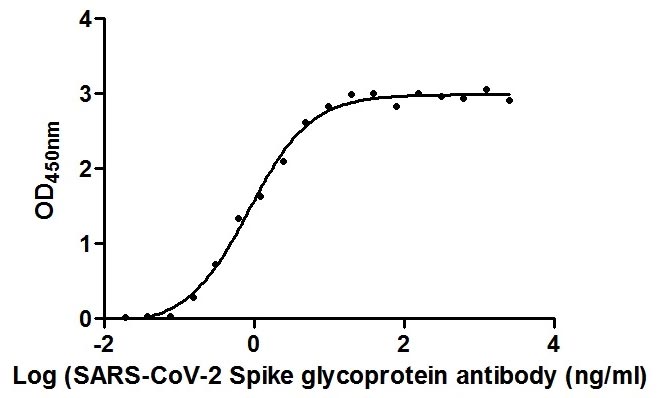
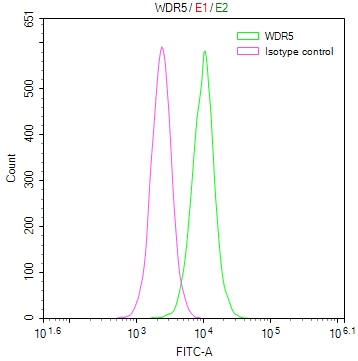
The Binding Activity of SARS-CoV-2 Spike RBD Nanobody with SARS-CoV-2-S1-RBD
Activity: Measured by its binding ability in a functional ELISA. Immobilized SARS-CoV-2-S1-RBD (CSB-YP3324GMY1) at 2 μg/ml can bind SARS-CoV-2 Spike RBD Nanobody, the EC50 is 0.8674 ng/ml. -
In the Colloidal Gold Immunochromatography Assay detection system, the background of antibody (CSB-RA33245A2GMY) is clean, the detection limit can be as low as 25ng/ml (1.75ng/0.07ml), and the sensitivity is very good.
-
SARS-CoV-2 Spike RBD Nanobody (CSB-RA33245A2GMY) competed with ACE2-HRP conjugate (CSB-MP866317HU) for binding to SARS-CoV-2-S1-RBD (CSB-YP3324GMY1). The binding signal of SARS-CoV-2-S1-RBD and ACE2-HRP conjugate was gradually reduced as the SARS-CoV-2 Spike RBD Nanobody concentrations increased. It indicated that this SARS-CoV-2 Spike RBD Nanobody effectively inhibited the SARS-CoV-2-S1-RBD/ACE2 binding. And the IC50 of this SARS-CoV-2 Spike RBD Nanobody is 1.296 nM.
-
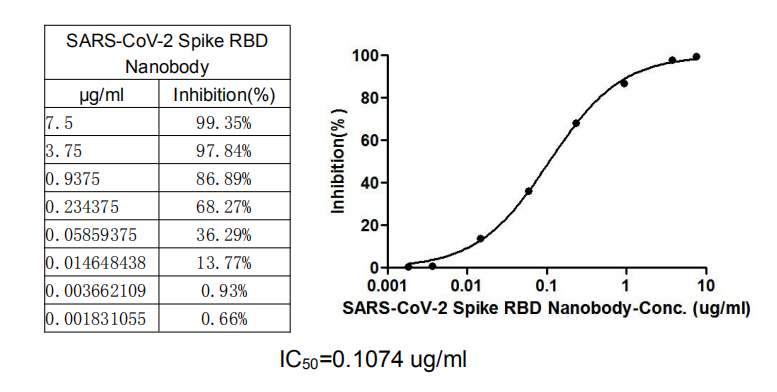
SARS-CoV-2 Spike RBD Nanobody (CSB-RA33245A2GMY) competitively prevented SARS-CoV-2-S1-RBD (CSB-YP3324GMY1) from binding to ACE2-HRP conjugate (CSB-MP866317HU). The inhibition efficacy of the SARS-CoV-2-S1-RBD/ACE2 binding was positively proportionally to the SARS-CoV-2 Spike RBD Nanobody concentrations. It showed that this SARS-CoV-2 Spike RBD Nanobody effectively inhibited the SARS-CoV-2-S1-RBD/ACE2 binding. And the IC50 of this SARS-CoV-2 Spike RBD Nanobody is 0.1074 μg/ml.
-
SARS-CoV-2 Spike protein RBD His/Sumostar Tag (CSB-YP3324GMY1) captured on COOH chip binding to the SARS-CoV-2 Spike RBD Nanobody (CSB-RA33245A2GMY), increases the local refractive index (RI), leading to a red shift of the LSPR peak position. The higher concentrations of SARS-CoV-2 Spike RBD Nanobody, the larger the wavelength shift. The detected affinity constant of SARS-CoV-2 Spike protein RBD/SARS-CoV-2 Spike RBD Nanobody binding is 28.2nM.
-
ELISA: Immobilize various types of SARS proteins at concentration of 2μg/ml on solid substrate, then react with SARS-CoV-2 Spike RBD Nanobody at concentration of 100μg/ml, 10μg/ml and 1μg/ml. It shows the SARS-CoV-2 Spike RBD Nanobody (CSB-RA33245A2GMY) is specific for SARS-CoV-2-S1-RBD protein, without any cross-reactivity with MERS-CoV, SARS-CoV, HCoV-OC43 or HCoV-229E.
-
-
其他:
產(chǎn)品詳情
-
產(chǎn)品描述:
This SARS-CoV-2 S1-RBD (Spike Glycoprotein S1 receptor-binding domain) antibody is a recombinant monoclonal antibody (also a Nanobody) generated through the expression of a DNA sequence inserting a human IgG1 Fc domain at the C-terminus, in human embryonic kidney 293 cells (HEK293). The DNA sequence encodes the SARS-CoV-2 spike RBD. The antibody is purified by protein G in vitro. It has been validated with high reactivity towards SARS-CoV-2 S1-RBD by a functional ELISA and good sensitivity for human SARS-CoV-2 spike glycoprotein (S protein) via the Colloidal Gold Immunochromatography Assay (GICA).
The SARS-CoV-2 S1-RBD Nanobody is also validated in Neutralizing and LSPR. In neutralizing assay, the binding signal of SARS-CoV-2 S1 RBD and ACE2 was inhibited by SARS-CoV-2 Spike RBD Nanobody. The IC50 is typically 0.1074 ug/ml. In the LSPR assay, the SARS-CoV-2 S1 RBD antibody showed a high affinity with SARS-CoV-2 Spike protein RBD (affinity constant: 28.2nM).
Specifically binding and recognizing the RBD of the SARS-CoV-2 spike glycoprotein, the SARS-CoV-2 S1 RBD antibody can react with samples infected with human coronavirus SARS-CoV-2. But it does not respond to MERS or SARS-CoV spike protein. Akin to other nanobodies, this recombinant nanobody is small and stable, which allows for its reaching to hidden epitopes such as crevices of target proteins.
-
Uniprot No.:
-
別名:S; 2; Spike glycoprotein; S glycoprotein; E2; Peplomer protein)
-
反應(yīng)種屬:Human Novel Coronavirus (SARS-CoV-2/ 2019-nCoV)
-
免疫原:Recombinant Human Novel Coronavirus Spike glycoprotein(S) (319-541aa) (CSB-YP3324GMY1 and CSB-MP3324GMY1b1)
-
免疫原種屬:Human Novel Coronavirus (SARS-CoV-2/ 2019-nCoV)
-
標記方式:Non-conjugated
-
克隆類型:Monoclonal
-
抗體亞型:VHH fusion with human IgG1 Fc
-
純化方式:Affinity-chromatography
-
克隆號:A1
-
濃度:It differs from different batches. Please contact us to confirm it.
-
保存緩沖液:Preservative: 0.03% Proclin 300
Constituents: 50% Glycerol, 0.01M PBS, pH 7.4 -
產(chǎn)品提供形式:Liquid
-
應(yīng)用范圍:ELISA, GICA, Neutralising
-
推薦稀釋比:
Application Recommended Dilution ELISA 1:10000-1:100000 GICA 1:10000-1:40000 Neutralising 1:100-1:10000 -
Protocols:
-
儲存條件:Upon receipt, store at -20°C or -80°C. Avoid repeated freeze.
-
貨期:Basically, we can dispatch the products out in 1-3 working days after receiving your orders. Delivery time maybe differs from different purchasing way or location, please kindly consult your local distributors for specific delivery time.
相關(guān)產(chǎn)品
靶點詳情
-
功能:attaches the virion to the cell membrane by interacting with host receptor, initiating the infection. Binding to human ACE2 receptor and internalization of the virus into the endosomes of the host cell induces conformational changes in the Spike glycoprotein. Binding to host NRP1 and NRP2 via C-terminal polybasic sequence enhances virion entry into host cell. This interaction may explain virus tropism of human olfactory epithelium cells, which express high level of NRP1 and NRP2 but low level of ACE2. The stalk domain of S contains three hinges, giving the head unexpected orientational freedom. Uses human TMPRSS2 for priming in human lung cells which is an essential step for viral entry. Can be alternatively processed by host furin. Proteolysis by cathepsin CTSL may unmask the fusion peptide of S2 and activate membranes fusion within endosomes.; mediates fusion of the virion and cellular membranes by acting as a class I viral fusion protein. Under the current model, the protein has at least three conformational states: pre-fusion native state, pre-hairpin intermediate state, and post-fusion hairpin state. During viral and target cell membrane fusion, the coiled coil regions (heptad repeats) assume a trimer-of-hairpins structure, positioning the fusion peptide in close proximity to the C-terminal region of the ectodomain. The formation of this structure appears to drive apposition and subsequent fusion of viral and target cell membranes.; Acts as a viral fusion peptide which is unmasked following S2 cleavage occurring upon virus endocytosis.; May down-regulate host tetherin (BST2) by lysosomal degradation, thereby counteracting its antiviral activity.
-
基因功能參考文獻:
- Study presents crystal structure of C-terminal domain of SARS-CoV-2 (SARS-CoV-2-CTD) spike S protein in complex with human ACE2 (hACE2); hACE2-binding mode similar overall to that observed for SARS-CoV. However, details at the binding interface show that key residue substitutions in SARS-CoV-2-CTD slightly strengthen the interaction and lead to higher affinity for receptor binding than SARS-CoV receptor-binding domain. PMID: 32378705
- crystal structure of the receptor-binding domain (RBD) of the spike protein of SARS-CoV-2 bound to the cell receptor ACE2 PMID: 32365751
- crystal structure of the receptor-binding domain (RBD) of the spike protein of SARS-CoV-2 (engineered to facilitate crystallization) in complex with ACE2 PMID: 32320687
- Out of the two isolates from India compared to the isolates from Wuhan, China, one was found to harbor a mutation in its receptor-binding domain (RBD) at position 407 where, arginine was replaced by isoleucine. This mutation has been seen to change the secondary structure of the protein at that region and this can potentially alter receptor binding of the virus. PMID: 32275855
- Structural modeling of the SARS-CoV-2 spike glycoprotein show similar receptor utilization between SARS-CoV-2 and SARS-CoV, despite a relatively low amino acid similarity in the receptor binding module. Compared to SARS-CoV and all other coronaviruses in Betacoronavirus lineage B, an extended structural loop containing basic amino acids were identified at the interface of the receptor binding (S1) and fusion (S2) domains. PMID: 32245784
- crystal structure of CR3022, a neutralizing antibody from a SARS patient, in complex with the receptor-binding domain of the SARS-CoV-2 spike (S) protein to 3.1 A; study provides insight into how SARS-CoV-2 can be targeted by the humoral immune response and revealed a conserved, but cryptic epitope shared between SARS-CoV-2 and SARS-CoV PMID: 32225176
- SARS-CoV and SARS-CoV-2 spike proteins have comparable binding affinities achieved by balancing energetics and dynamics. The SARS-CoV-2-ACE2 complex contains a higher number of contacts, a larger interface area, and decreased interface residue fluctuations relative to the SARS-CoV-ACE2 complex. PMID: 32225175
- Interaction interface between cat/dog/pangolin/Chinese hamster ACE2 and SARS-CoV/SARS-CoV-2 S protein was simulated through homology modeling. Authors identified that N82 of ACE2 showed closer contact with receptor-binding domain of S protein than human ACE2. PMID: 32221306
- SARS-CoV-2 S glycoprotein harbors a furin cleavage site at the boundary between the S1/S2 subunits, which is processed during biogenesis and sets this virus apart from SARS-CoV and SARS-related CoVs; determined cryo-EM structures of the SARS-CoV-2 S ectodomain trimer. PMID: 32201080
- Study demonstrates that SARS-CoV-2 uses the SARS-CoV receptor ACE2 for entry and the serine protease TMPRSS2 for S protein priming. PMID: 32155444
- The ACE2-B0AT1 complex exists as a dimer of heterodimers. Structural alignment of the RBD-ACE2-B0AT1 ternary complex with the S protein of SARS-CoV-2 suggests that two S protein trimers can simultaneously bind to an ACE2 homodimer. PMID: 32142651
- study demonstrated SARS-CoV-2 S protein entry on 293/hACE2 cells is mainly mediated through endocytosis, and PIKfyve, TPC2 and cathepsin L are critical for virus entry; found that SARS-CoV-2 S protein could trigger syncytia in 293/hACE2 cells independent of exogenous protease; there was limited cross-neutralization activity between convalescent sera from SARS and COVID-19 patients PMID: 32132184
- study determined a 3.5-angstrom-resolution cryo-electron microscopy structure of the 2019-nCoV S trimer in the prefusion conformation; provided biophysical and structural evidence that the 2019-nCoV S protein binds angiotensin-converting enzyme 2 (ACE2) with higher affinity than does severe acute respiratory syndrome (SARS)-CoV S PMID: 32075877
顯示更多
收起更多
-
亞細胞定位:Virion membrane; Single-pass type I membrane protein. Host endoplasmic reticulum-Golgi intermediate compartment membrane; Single-pass type I membrane protein. Host cell membrane; Single-pass type I membrane protein.
-
蛋白家族:Betacoronaviruses spike protein family
Most popular with customers
-
-
YWHAB Recombinant Monoclonal Antibody
Applications: ELISA, WB, IF, FC
Species Reactivity: Human, Mouse, Rat
-
-
-
-
-
-